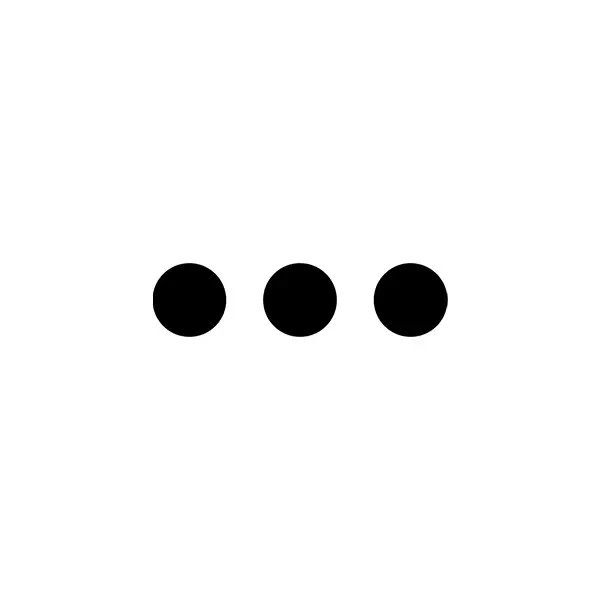
By exploiting our robust Synthetic Genomics platform, we reduce the time to market by completely bypassing the major strain development bottlenecks.
We have developed an entirely new technology platform for Synthetic Genomics that enables us to develop customized strains suited for a diverse range of applications. Starting from the desired end uses for a strain, we design and assemble customized secondary genomes that are uniquely purpose-built. These rationally designed genomes encode genes and complete biosynthetic pathways that enable the cell to perform specific functions. This is a disruptive technology that can be suitable to a wide-range of potential applications and uses.
For more than 10 years, our research has focused on the impact that chromosomal folding and DNA looping have on gene expression. We have found that the expression of a given gene can vary >1000x fold depending solely upon where the gene is positioned within the chromosome as well as other factors (local context, orientation, presence of enzymes and DNA binding proteins). We have built computational tools that enable us to predict and analyze how a gene will “behave” in a certain chromosomal context. We apply these natural rules into the design of optimized expression systems, namely secondary chromosomes. These expression vectors are purpose-built to produce high-added value products.

Our technology platform consists of a suite of proprietary tools, both computational and molecular, for the design and assembly of genomes. In addition to the new tools that we have invented, we have established a computer guided genome development pipeline that streamlines the engineering process. This pipeline begins by using proprietary algorithms to analyse relevant genomic datasets, followed by the in silico design of the genome. The DNA fragments to be purchased are computationally optimised to increase the success rate of DNA synthesis and ordered from a company. We then assemble the fragments into a functional genome using a variety of proprietary in vitro and in vivo methods.
Our proprietary genome editing methods are now commercially available in the form of the Simply Seamless DNA assembly and cloning kit.
The “classical” strain development approach consists of inserting a gene of interest into a micro-organism’s chromosome, at a random position. This is then repeated thousands of times at different positions throughout the organism’s genome resulting in a library of thousands of different strains. The performance of each strain is then assessed to identify the most promising candidate strains (shot-gun random approach). This process is laborious, costly and time-consuming.
Synovance’s Synthetic Genomic Technology uses our knowledge of the fundamental parameters that enhance gene expression along with our proprietary genomic tools for identifying the genetic context to design a genome that will result in robust strain performance (3D chromosome conformation, local genomic context, gene orientation, etc). The optimized strain is then engineered based upon these design parameters. This results in a high-performance strain in terms of gene expression, typically on the first attempt with no de-bugging required (bottom-up rational approach). Our process is extremely focused, cost-effective and rapid.